Filtering
Warning in as.POSIXlt.POSIXct(Sys.time()): unknown timezone 'zone/tz/2018c.
1.0/zoneinfo/America/Los_Angeles'Last updated: 2018-04-23
Code version: 90feb3f
Summary
We imported raw data (nasal_raw.rds) and excluded samples and OTUs based on the following criteria. The filtered data are saved in nasal_filtered.rds.
Prior to filtering, there are 42,999 features and 201 samples
- Low sequencing depth
1.1. Samples with total count < 1000
1.2. OTUs not present (> 0 read) in any sample
1.3. Satisfies 1.1 AND 1.2
After Step 1, left with 42,982 features and 197 samples.
- OTUs not present
2.1. Present (> 0 read) in 5 or more samples in 1
2.2. Present and quantified with > 20 read in at least one sample
2.3. Satisfies 2.1 OR 2.2, otherwise excluded.
After Step 2, left with 5,521 features and 197 samples.
- Excluded OTUs not assigned at the Kindom level
After Step 3, left with 4,423 features and 197 samples
In the end, 4 samples/patients were excluded from the analysis. See the bottom of this page for sample demographics.
\(~\)
Questions:
Maybe included one patient that has ~900 total reads?
Difference between treatment and control demographics before/after excluded these samples?
Packages
library(metagenomeSeq)Loading data
MRobj = readRDS("../data/nasal_raw.rds")
MRobjMRexperiment (storageMode: environment)
assayData: 42999 features, 201 samples
element names: counts
protocolData: none
phenoData
sampleNames: EM0042 EM0047 ... E0168 (201 total)
varLabels: StudyID age ... infnone (94 total)
varMetadata: labelDescription
featureData
featureNames: denovo0 denovo1 ... denovo42998 (42999 total)
fvarLabels: Kingdom Phylum ... Species (7 total)
fvarMetadata: labelDescription
experimentData: use 'experimentData(object)'
Annotation: \(~\)
Filter samples with low depth
Filter out samples with total count < 1000.
Filter out OTUs with total count across samples < 1. In other words, present in at least one sample.
hist(log2(colSums(MRobj)),xlab="Log sequencing depth",
main = "Log sequencing depth")
abline(v=log2(1000))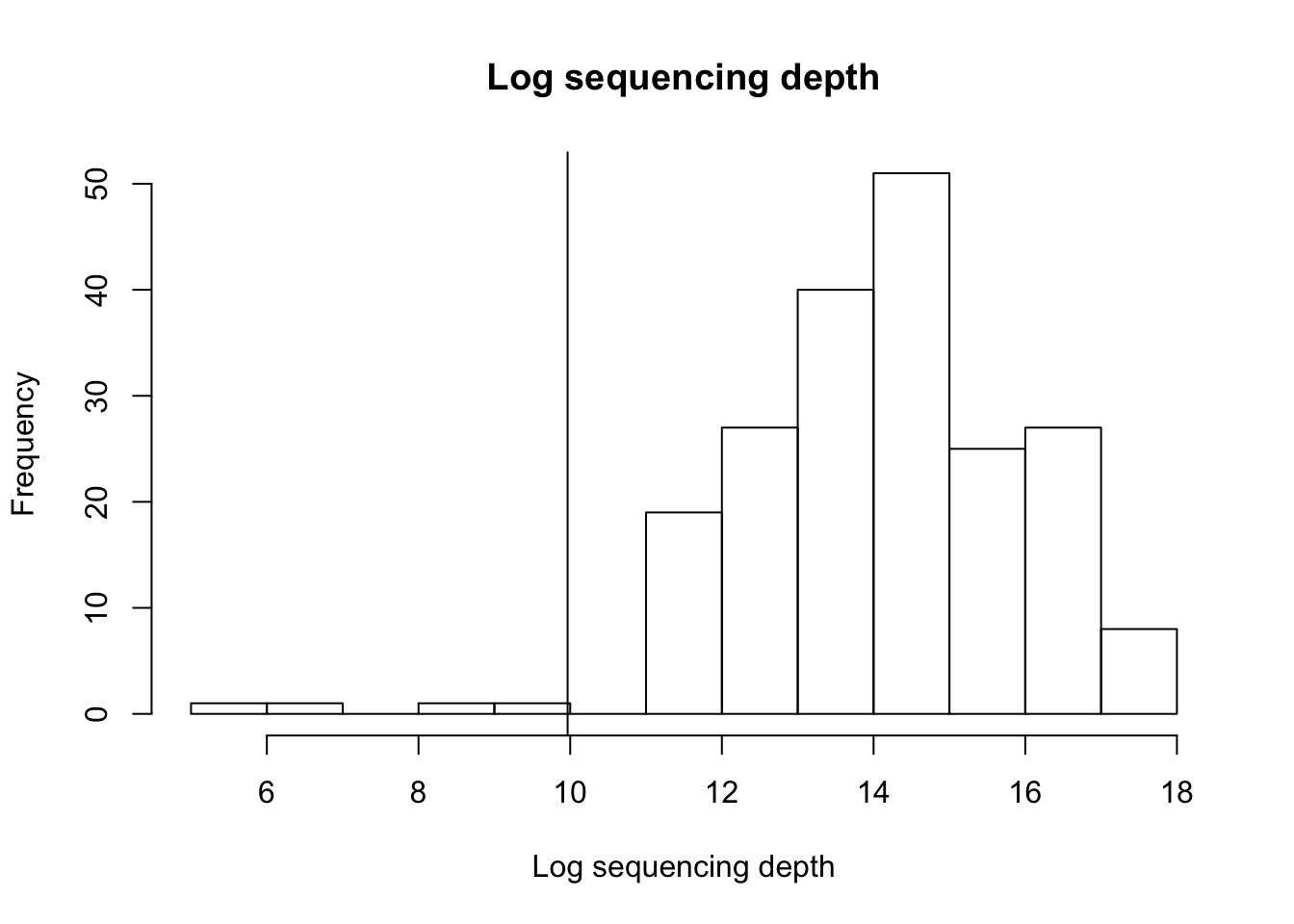
MRobj = filterData(MRobj, depth=1000, present=1)
MRobjMRexperiment (storageMode: environment)
assayData: 42982 features, 197 samples
element names: counts
protocolData: none
phenoData
sampleNames: EM0042 EM0047 ... E0168 (197 total)
varLabels: StudyID age ... infnone (94 total)
varMetadata: labelDescription
featureData
featureNames: denovo0 denovo1 ... denovo42998 (42982 total)
fvarLabels: Kingdom Phylum ... Species (7 total)
fvarMetadata: labelDescription
experimentData: use 'experimentData(object)'
Annotation: \(~\)
Filter OTUs not present
OTUs present (> 0 read) in 5 or more samples
OTUs that are quantified with > 20 reads in at least one sample
After filtering, we have 5,521 features and 197 samples.
nmat = MRcounts(MRobj)
keep = which( rowSums(nmat>0)>=5 | rowSums(nmat>=20)>0 )
MRobj=MRobj[keep,]
MRobjMRexperiment (storageMode: environment)
assayData: 5521 features, 197 samples
element names: counts
protocolData: none
phenoData
sampleNames: EM0042 EM0047 ... E0168 (197 total)
varLabels: StudyID age ... infnone (94 total)
varMetadata: labelDescription
featureData
featureNames: denovo2 denovo6 ... denovo42992 (5521 total)
fvarLabels: Kingdom Phylum ... Species (7 total)
fvarMetadata: labelDescription
experimentData: use 'experimentData(object)'
Annotation: Unassigned
Number of unique organisms we find within each clade. PS. 2 at the Kindom level due to the unassigned
sapply(fData(MRobj),function(i)length(unique(i)))Kingdom Phylum Class Order Family Genus Species
2 31 75 144 259 478 554 Number of features unassigned at the Kindom level. (1,098 out of total 5521 OTUs)
length(which(fData(MRobj)[,1]=="Unassigned"))[1] 1098Filtering Unassigned.
MRobj = MRobj[- which(fData(MRobj)[,1]=="Unassigned"),]
saveRDS(MRobj,file="../data/nasal_filtered.rds")After filtering, we have 4,423 features and 197 samples.
MRobjMRexperiment (storageMode: environment)
assayData: 4423 features, 197 samples
element names: counts
protocolData: none
phenoData
sampleNames: EM0042 EM0047 ... E0168 (197 total)
varLabels: StudyID age ... infnone (94 total)
varMetadata: labelDescription
featureData
featureNames: denovo6 denovo17 ... denovo42992 (4423 total)
fvarLabels: Kingdom Phylum ... Species (7 total)
fvarMetadata: labelDescription
experimentData: use 'experimentData(object)'
Annotation: Excluded sample
4 samples were excluded from the filtered data: EM0062, E0194, EM0088, EM0053.
pre_filtered <- readRDS("../data/nasal_raw.rds")
post_filtered <- readRDS("../data/nasal_filtered.rds")
excluded_samples <- setdiff(colnames(MRcounts(pre_filtered)), colnames(MRcounts(post_filtered)))
excluded_samples[1] "EM0062" "E0194" "EM0088" "EM0053"colSums(MRcounts(pre_filtered))[colnames(pre_filtered) %in% c("EM0062", "E0194", "EM0088", "EM0053")]EM0062 E0194 EM0088 EM0053
104 934 271 36 Phenotypes of the 4 excluded samples.
EM0062
Demographics: age 66, male, white, underwent cardiac surgery, studysite = 1, seqrun = 2
Primary outcome: anyinf6m (infection 6 mo. post surgery) = 1, naswabsa1 (nasal swab culture positive) = 1
pData(pre_filtered)[rownames(pData(pre_filtered)) == "EM0062",] StudyID age gender race srgtpop htprop wtprop bmiprop ster3prop
EM0062 EM0062 66 0 0 1 180.3 114 35.1 0
cadprop chfdgprop htnprop cholprop dbprop smokcuprop cvdprop
EM0062 1 1 1 0 1 0 0
padprop lungdisprop asthmaprop dialprop cancerprop obesity
EM0062 0 0 0 0 1 1
creatprop bunprop hgbprop hctprop wbcprop plateprop wbc hgb hct
EM0062 0.91 17 13.7 39.9 6600 237 6400 15.7 46.9
plate ssi ssidepth SSIorg1 SSIorg2 SSIorg3 infdeath cardsrgsc
EM0062 191 1 deep none <NA> <NA> 0 1
cransrgsc spinsrgsc vascsrgsc anyinf30 anyinf6m death6m ssideep6m
EM0062 0 0 0 1 1 0 1
bloodinf6m pneumon6m naswabtp studysite DateExtracted seqrun
EM0062 0 0 3 1 2014-05-12 2
infabprop miprop smokprprop ulcerprop liverprop readm_30d maristat3
EM0062 0 0 1 0 0 1 1
naswabsa1 naswabmrsa1 insurstat3 CharlCA Charlcat educat4
EM0062 1 0 2 0 1 1
nonsysester1prop bronmedprop lungdz medincome hosprop asa CharlF
EM0062 0 0 0 56221 1 4 2
COPD h2blockprop skinev1prop nasalster antihist allergies outinop
EM0062 0 1 0 0 0 0 1
dtop ssideepsepi ssideepSA ssideepStaph ssideepstrep
EM0062 09/04/2009 0 0 0 0
ssideeppseudo ssideepentero ssideepnone infSepi infSA infStaph
EM0062 0 0 1 0 0 0
infStrep infpseudo infEntero infnone
EM0062 0 0 0 1\(~\)
E0194
Demographics: age 66, male, white, underwent cardiac surgery, study site = 0, seqrun = 2
Primary outcome: anyinf6m (infection 6 mo. post surgery) = 0, naswabsa1 (nasal swab culture positive) = 0
pData(pre_filtered)[rownames(pData(pre_filtered)) == "E0194",] StudyID age gender race srgtpop htprop wtprop bmiprop ster3prop
E0194 E0194 66 0 0 1 165 88 32.3 0
cadprop chfdgprop htnprop cholprop dbprop smokcuprop cvdprop padprop
E0194 1 0 1 1 0 0 0 0
lungdisprop asthmaprop dialprop cancerprop obesity creatprop bunprop
E0194 0 0 1 0 1 4.9 28
hgbprop hctprop wbcprop plateprop wbc hgb hct plate ssi ssidepth
E0194 8.8 28.4 3170 207 2600 8.1 25.6 201 0 <NA>
SSIorg1 SSIorg2 SSIorg3 infdeath cardsrgsc cransrgsc spinsrgsc
E0194 <NA> <NA> <NA> 1 1 0 0
vascsrgsc anyinf30 anyinf6m death6m ssideep6m bloodinf6m pneumon6m
E0194 0 0 0 1 0 0 0
naswabtp studysite DateExtracted seqrun infabprop miprop smokprprop
E0194 2 0 2013-08-21 2 1 1 1
ulcerprop liverprop readm_30d maristat3 naswabsa1 naswabmrsa1
E0194 0 0 0 1 0 0
insurstat3 CharlCA Charlcat educat4 nonsysester1prop bronmedprop
E0194 1 0 3 0 0 0
lungdz medincome hosprop asa CharlF COPD h2blockprop skinev1prop
E0194 0 41580 1 4 5 0 0 1
nasalster antihist allergies outinop dtop ssideepsepi
E0194 0 0 0 2 02/20/2008 0
ssideepSA ssideepStaph ssideepstrep ssideeppseudo ssideepentero
E0194 0 0 0 0 0
ssideepnone infSepi infSA infStaph infStrep infpseudo infEntero
E0194 0 0 0 0 0 0 0
infnone
E0194 0\(~\)
EM0088
Demographics: age 74, female, white, underwent cardiac surgery, studysite = 1, seqrun = 2
Primary outcome: anyinf6m (infection 6 mo. post surgery) = 1, naswabsa1 (nasal swab culture positive) = 1
pData(pre_filtered)[rownames(pData(pre_filtered)) == "EM0088",] StudyID age gender race srgtpop htprop wtprop bmiprop ster3prop
EM0088 EM0088 74 1 0 1 160 65.4 25.5 0
cadprop chfdgprop htnprop cholprop dbprop smokcuprop cvdprop
EM0088 0 0 1 0 0 1 0
padprop lungdisprop asthmaprop dialprop cancerprop obesity
EM0088 0 0 0 0 0 0
creatprop bunprop hgbprop hctprop wbcprop plateprop wbc hgb hct
EM0088 1.73 34 12.5 36.7 6300 156 6000 12.6 36.5
plate ssi ssidepth SSIorg1 SSIorg2 SSIorg3 infdeath
EM0088 156 1 deep enterococcus bacteroides pseudomonas 0
cardsrgsc cransrgsc spinsrgsc vascsrgsc anyinf30 anyinf6m death6m
EM0088 1 0 0 0 1 1 1
ssideep6m bloodinf6m pneumon6m naswabtp studysite DateExtracted
EM0088 1 1 1 3 1 2014-05-12
seqrun infabprop miprop smokprprop ulcerprop liverprop readm_30d
EM0088 2 1 0 1 0 0 0
maristat3 naswabsa1 naswabmrsa1 insurstat3 CharlCA Charlcat educat4
EM0088 1 1 1 1 0 3 0
nonsysester1prop bronmedprop lungdz medincome hosprop asa CharlF
EM0088 0 1 1 95094 1 4 5
COPD h2blockprop skinev1prop nasalster antihist allergies outinop
EM0088 1 1 0 0 0 0 1
dtop ssideepsepi ssideepSA ssideepStaph ssideepstrep
EM0088 07/13/2011 0 0 0 1
ssideeppseudo ssideepentero ssideepnone infSepi infSA infStaph
EM0088 1 0 0 0 0 0
infStrep infpseudo infEntero infnone
EM0088 1 1 0 0\(~\)
EM0053
Demographics: age 64, male, white, underwent cardiac surgery, studysite = 1, seqrun = 2
Primary outcome: anyinf6m (infection 6 mo. post surgery) = 1, naswabsa1 (nasal swab culture positive) = 0
pData(pre_filtered)[rownames(pData(pre_filtered)) == "EM0053",] StudyID age gender race srgtpop htprop wtprop bmiprop ster3prop
EM0053 EM0053 64 0 0 1 177.8 94 29.7 0
cadprop chfdgprop htnprop cholprop dbprop smokcuprop cvdprop
EM0053 1 0 1 0 0 0 0
padprop lungdisprop asthmaprop dialprop cancerprop obesity
EM0053 0 0 0 0 0 0
creatprop bunprop hgbprop hctprop wbcprop plateprop wbc hgb hct
EM0053 1.17 8 15.5 42.4 5900 202 4200 13.7 38
plate ssi ssidepth SSIorg1 SSIorg2 SSIorg3 infdeath cardsrgsc
EM0053 5 1 deep klebsiella proteus <NA> 0 1
cransrgsc spinsrgsc vascsrgsc anyinf30 anyinf6m death6m ssideep6m
EM0053 0 0 0 1 1 0 1
bloodinf6m pneumon6m naswabtp studysite DateExtracted seqrun
EM0053 1 1 3 1 2014-05-12 2
infabprop miprop smokprprop ulcerprop liverprop readm_30d maristat3
EM0053 1 1 0 0 0 0 0
naswabsa1 naswabmrsa1 insurstat3 CharlCA Charlcat educat4
EM0053 0 0 2 0 1 1
nonsysester1prop bronmedprop lungdz medincome hosprop asa CharlF
EM0053 0 0 0 92509 0 4 1
COPD h2blockprop skinev1prop nasalster antihist allergies outinop
EM0053 0 1 1 0 0 0 1
dtop ssideepsepi ssideepSA ssideepStaph ssideepstrep
EM0053 07/14/2009 0 0 0 0
ssideeppseudo ssideepentero ssideepnone infSepi infSA infStaph
EM0053 0 1 0 0 0 0
infStrep infpseudo infEntero infnone
EM0053 0 0 1 0Session Information
R version 3.4.1 (2017-06-30)
Platform: x86_64-apple-darwin15.6.0 (64-bit)
Running under: macOS High Sierra 10.13
Matrix products: default
BLAS: /Library/Frameworks/R.framework/Versions/3.4/Resources/lib/libRblas.0.dylib
LAPACK: /Library/Frameworks/R.framework/Versions/3.4/Resources/lib/libRlapack.dylib
locale:
[1] en_US.UTF-8/en_US.UTF-8/en_US.UTF-8/C/en_US.UTF-8/en_US.UTF-8
attached base packages:
[1] parallel stats graphics grDevices utils datasets methods
[8] base
other attached packages:
[1] metagenomeSeq_1.20.1 RColorBrewer_1.1-2 glmnet_2.0-13
[4] foreach_1.4.4 Matrix_1.2-12 limma_3.34.9
[7] Biobase_2.38.0 BiocGenerics_0.24.0
loaded via a namespace (and not attached):
[1] Rcpp_0.12.15 knitr_1.20 magrittr_1.5
[4] lattice_0.20-35 stringr_1.3.0 caTools_1.17.1
[7] tools_3.4.1 grid_3.4.1 KernSmooth_2.23-15
[10] git2r_0.21.0 gtools_3.5.0 htmltools_0.3.6
[13] iterators_1.0.9 matrixStats_0.53.1 yaml_2.1.18
[16] rprojroot_1.3-2 digest_0.6.15 bitops_1.0-6
[19] codetools_0.2-15 evaluate_0.10.1 rmarkdown_1.9
[22] gdata_2.18.0 stringi_1.1.6 compiler_3.4.1
[25] gplots_3.0.1 backports_1.1.2 This R Markdown site was created with workflowr